Projekt som vi stöttar
Projekt som vi stöttar
Medtech4Health är ett nationellt strategiskt Innovationsprogram som startade 2016 och är finansierat av VINNOVA.
På den här sidan kan du läsa projektsammanfattningar och lägesrapporter från de projekt som vi stöttar på något sätt. De flesta av projekten fått medel från oss via öppna utlysningar men även andra sätt kan förekomma.
Vi driver även interna aktiviteter i projektform, dessa hittar du i huvudmenyn.
Senaste nytt om och från projekt
Är ni ett medicintekniskt företag och behöver extern kompetens?
Idag öppnar utlysningen Kompetensförstärkning i småföretag som ger SMF en möjlighet att stärka kompetensen genom att anlita en extern person under en projektperiod. Syftet är att förkorta tiden från idé till patientnytta
SmartPAN vid Bukspottkörtelcancer närmar sig lansering med stöd av finansiering från Kompetensförstärkning i småföretag
MAGLE Chemoswed AB har utvecklat SmartPAN, en preventiv metod vid bukspottkörtelcancer. Genom medel från utlysningen Kompetensförstärkning i företag kan de nu ta stöd från expertis i arbetet med att marknadsintroducera produkten.
Regelefterlevnad – ett seminarium för dig inom innovationsstödsystemen
Kan regelefterlevnad vara en konkurrensfördel för forskare, små och medelstora företag inom medicinteknik? Och hur kan vi som jobbar i innovationsstödsystemet bidra? Välkommen till ett seminarium den 22 februari där vi redogör för den ökande regleringstakten inom medicinteknik och hälso- och sjukvård, samt förslag på hanteringssystem och utbildning.
Se inspelningen från webbinarium om Internationalisering i medicintekniska småföretag
Nu finns vårt webbinarium om internationalisering på Youtube. Vid två webbinarier presenterades resultatet av ett flera år långt arbete i vårt strategiska projekt Internationaliseringsprojektet i form av en spelplan. Presenterade gjorde Robert Harju-Jeanty.
Positiva resultat när världens första textila strålskyddskläder för operationspersonal utvärderades med hjälp av Kompetensförstärkningsmedel
Göteborgsbaserade företaget Texray tog chansen att utvärdera sina innovativa strålskyddskläder för operationspersonal kliniskt med hjälp av medel ur vår kompetensförstärkningsutlysning. Resultaten var mycket positiva- både vad gäller skyddsnivån och kundnöjdhet.
Kompetensförstärkningsmedel gav kvalitetsledningssystem för innovativa kompressionsprodukter
Finansiering från Kompetensförstärkningsutlysning för småföretag hjälpte det västsvenska företaget PressCise att implementera kvalitetsledningssystem för sina innovativa kompressionsprodukter.
Resultatdialog för Nationellt utvecklingsprojekt den 14–15 januari
Torsdagen och fredagen den 14 och 15 januari, från lunch till lunch, samlades delprojektledarna i Medtech4Healths Nationella utvecklingsprojekt för implementering av medicintekniska innovationer och tjänster (NUP) som har pågått under två år. Deltagarna presenterade och [...]
Real Heart som utvecklar världens första artificiella fyrkammarhjärta får kompetensförstärkningsmedel
Real Heart som utvecklar världens första artificiella fyrkammarhjärta får kompetensförstärkningsmedel Varje år dör cirka två miljoner människor i världen av hjärtsvikt. Bara i Sverige är det dödsorsaken för tio personer varje dag. Det [...]
AndningMed får Kompetensförstärkningsmedel för att planera klinisk studie av sin förbättring av inhalationsteknik
Start-up som utvecklar en produkt för att minska fel vid användning av inhalatorer får medel ur Kompetensförstärkningsutlysningen för att ta in professionell hjälp vid planering av klinisk studie.
Elsa Science får Kompetensförstärkningsmedel för att ta sin tjänst till USA
Elsa Science har nyligen beviljats medel ur vår utlysning Kompetensförstärkning. Medlen ska användas till att förstärka kompetensen vad gäller regulatoriskt arbete inför en lansering på USA-marknaden.
Rekryteringstips för start-ups – se vår Innovation Hive med Åsa Glavich på Youtube
Rekryteringstips för start-ups - se vår Innovation Hive med Åsa Glavich på Youtube Den 28 oktober arrangerade vi tillsammans med vår systerorganisation Swelife ännu en Innovation Hive- denna gång med fokus på rekrytering. [...]
Samverkan A och O för den funktionella proteshylsan
Christoffer Lindhe lever med tre proteser och driver företaget Lindhe Xtend vilket finns representerat på en global marknad. Han ser dagligen många möjliga förbättringar som kan göras av proteser och sökte upp Linda Nyden, projektkoordinator på Smart Textiles, Högskolan i Borås, för att diskutera sin idé närmare.
Watch Isabel Gonçalves presentation from the Inclusive Innovation seminar
Watch Isabel Gonçalves presentation from the seminar held on 31st August as a part of Inclusive Innovation's launching of the new work material.
Resultaten från Inkluderande Innovation presenteras i Stockholm och i norr
Medtech4Health har under de senaste åren jobbat med inkluderingsfrågor i projektet Inkluderande Innovation. Under senhösten kommer du att få möjlighet att ta del av projektresultaten och få konkreta verktyg för arbetet. Få inspiration och ställ frågor om materialet direkt till projektledare Malin Hollmark och inkluderingsexperten Pernilla Alexandersson från AddGender.
Torbjörn Kronander ger expertråd om försäljning på Innovation Hive
På höstens första Innovation Hive fick vi besök av Torbjörn Kronander, VD och grundare till den medicintekniska delen på Sectra, och också styrelseledamot i Medtech4Health. Torbjörn har en lång och värdefull erfarenhet av att bygga upp medicintekniska bolag från grunden och att ta dem ut på den internationella marknaden och på detta seminarium bad vi Torbjörn att ge råd om hur man kan tänka för att bygga upp försäljningsteam och utveckla försäljningen för att lyckas.
Rundabordssamtal i Inkluderande innovation under Medicinteknikdagarna 2021
Under Medicinteknikdagarna 2021 arrangerar vårt strategiska projekt Inkluderande Innovation ett rundabordssamtal med tanken att vi tillsammans ska lyfta viktiga frågeställningar och goda exempel för att skapa ännu starkare och mer konkurrenskraftiga svenska innovationsföretag.
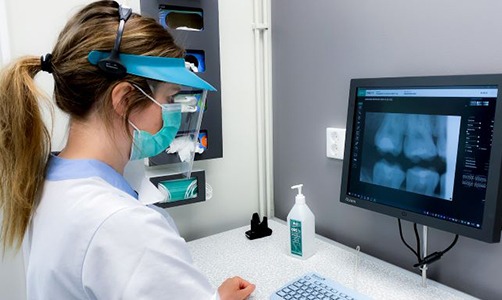
Innovationsmotor i Västerbotten tillvaratar värdefulla tandvårdsresurser
Innovationsmotor i Västerbotten tillvaratar värdefulla tandvårdsresurser Innovationsmotorer är ett strategiskt projekt som finansieras av Medtech4Health. Projektet leds nationellt av Lena Svendsen på Swedish Medtech, en av Medtech4Healths noder. Hej Lena, vad gör man [...]
Inspelning från Inkluderande Innovations lanseringsevent nu på Youtube
Om du missade de två lanseringsevent som det strategiska projektet Inkluderande Innovation höll den 30 och 31 augusti - varav ett på engelska - kan du nu se dem i efterhand textade på vår Youtube-kanal.
Se inspelning från seminariet om Horisont Europe med Nadine Schweizer
Nu finns inspelningen från seminariet om Horisont Europe med Nadine Schweizer i augusti 2021 att se på vår Youtube-kanal.
Medel för kompetensförstärkning bidrog till FDA-godkännande för cancerinnovation
Micropos senaste produkt har i september blivit FDA-godkänd med hjälp av finansiering från vår utlysning för Kompetensförstärkning i småföretag. Läs artikeln om hur medlen har bidragit i arbetet.
Diabetes-appen som kom till när dietistens och patient-representantens vägar korsades
Under våren korades vinnaren av Medtech4Health Innovation Award och ett av de tre finalistbidragen var T1D-appen. Vi har pratat med grundarna Elin Cederbrant och Elisabeth Jelleryd.
Seminarium om Horisont Europa – EU:s ramprogram för forskning och innovation 2021-2027
På detta seminarium om Horisont Europa - EU:s ramprogram för forskning och innovation 2021-2027 introducerar Nadine Schweizer, National contact point, Horizon Europe, Vinnova, oss till Horisont Europas hälsosatsningar där medicinsktekniska lösningar ingår. Därefter blir det plats för frågor och diskussion.
Vad händer i fokusområde systematisk utvärdering – ett delprojekt inom Nationella utvecklingsprojektet för implementering (NUP)?
Vi pratar med Adam Darwich, projektledare för fokusområde systematisk utvärdering – ett delprojekt inom Medtech4Healths strategiska projekt Nationella utvecklingsprojektet för implementering (NUP).
Vi lanserar material för att arbeta mer jämställt och inkluderande i medicintekniska projekt och verksamheter
I höst lanserar vi vårt material för att hjälpa dig att arbeta jämställt och inkluderande. Du kan välja mellan två lunchevent då vi presenterar materialet. Nu kan du anmäla dig!
Se prisutdelningsfilmen från Innovation Award 2021
Nu kan du se filmen från prisutdelningen för Innovation Award 2021 här!
Världsledande innovation resultatet när neurokirurger driver utvecklingen
Våren 2021 delades Medtech4Health Innovation Award ut till en världsledande forskargrupp bestående av fyra neurokirurger på Karolinska universitetssjukhuset. Det är Adrian Elmi Terander, Erik Edström, Gustav Burström och Oscar Persson som utvecklat en helt ny navigationsteknik för att med hjälp av Augmented Reality (AR) kunna se exakt var i ryggraden man befinner sig när man opererar.
Evira – ett digitalt verktyg för behandling av barnfetma
Evira – ett digitalt verktyg för behandling av barnfetma Hej Love Marcus på Evira. Ni har fått stöd ur två av våra utlysningar de senaste åren. Dels ur Kompetensförstärkning för små företag och [...]
Vad händer i projektet Inkluderande Innovation?
Vad händer i projektet Inkluderande Innovation? Läs intervjun med Malin Hollmark, projektledare.
Antimicrobial resistance the subject of the first InfraLife workshop
The first InfraLife workshop on 2 June gathers stakeholders in the field of antimicrobial resistance (AMR) for a joint discussion. This workshop is organised by SciLifeLab, MAXIV, ESS, SwedenBIO, LIF, SWElife, and MedTech4Health.
Zenicor fick medel ur Kompetensförstärkningsutlysningen
Nu har det beslutats vilka som får medel ur årets första avläsning i vår utlysning Kompetensförstärkning för småföretag. Det är Stockholmsbaserade Zenicor Medical Systems AB som får finansiering för MDR-anpassning av system för digitaliserad arytmidiagnostik.
SENASTE NYTT
Finansieringsglapp lämnar små och medelstora företag utan stöd
Medtech4Health och Swelife har tillsammans finansierat över 200 projekt genom utlysningen ”Samverkansprojekt för bättre hälsa”. Syftet har varit att accelerera utveckling och kommersialisering av innovativa lösningar för vård och hälsa. Resultatet talar sitt tydliga språk – tillväxten hos de finansierade företagen har ökat kraftigt.
Happy Summer Wishes from Medtech4Health
After an intense spring, it's almost time for a well-deserved summer break. But first, we look forward to meeting you in Almedalen, where we are part of the Tillsammans meeting place.
Medtech4Health on Tour: MedTech Forum Comes to Stockholm in 2026
This year, The MedTech Forum 2025 was held in Lisbon. Around 1,000 participants gathered at Europe’s largest health and medical technology conference for three days of insightful discussions on the sustainable future of healthcare in Europe.